
MEDIZINISCHE KLINIK UND POLIKLINIK I
Gastroenterologie, Hepatologie, Infektiologie, Nephrologie, Endokrinologie, Diabetologie
UNIVERSITÄTSKLINIKUM BONN
Prof. Dr. Christian Strassburg

 |
MEDIZINISCHE KLINIK UND POLIKLINIK I |
 |
 |
Wissenschaftliche StudienResearch Group "Systems Control of Metabolic Diseases"
|
|||||||||||||
|
Wichtige Telefonnummern
Patientenübernahme
Gastroenterologischer Notfall
Nephrologischer Notfall
Dienstarzt Innere Medizin
Konsiliardienste
Pforte Schnellzugriff Transplantationsbüro Sitemap COVID-19 Infos |
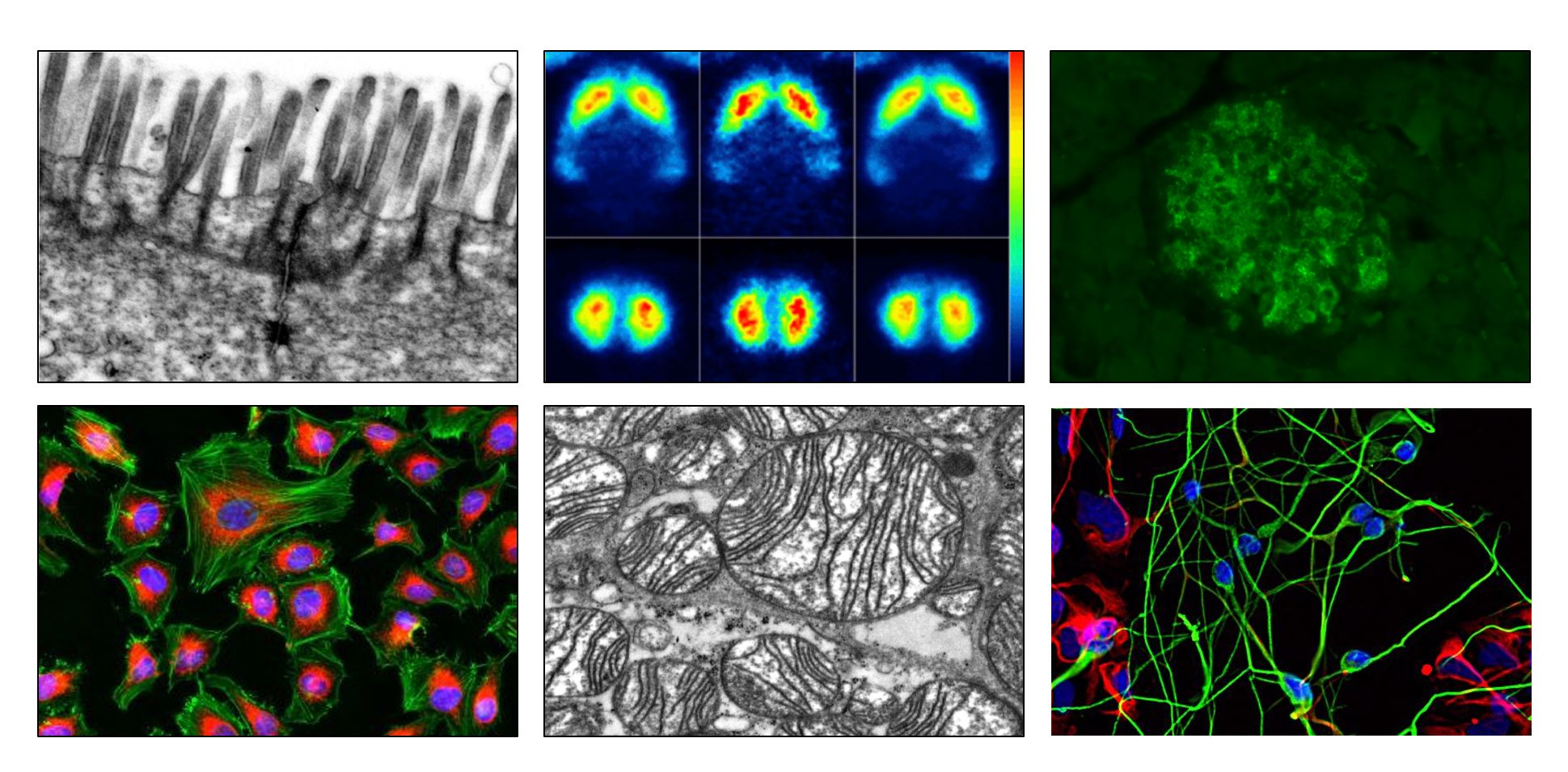
Top, left to right: REM image of intestinal epithelial cells, brain PET images, insulin staining of pancreatic islets. Bottom, left to right: fluorescence staining of subcellular compartments, mitochondria in brown adipose tissue, human stem cell derived progenitor cells and neurons (courtesy by the Institute of Reconstructive Neurobiology, UKB). RESEARCH INTERESTS:For the first time in history, a situation is reached where our planet harbors more obese than underweight inhabitants. Obesity and related metabolic diseases have reached pandemic proportions over the last few decades and will probably place an ever-increasing burden on our global health systems. The evolution of our modern environment with ready accessibility to palatable, energy-dense foods in virtually unlimited quantity combined with a dominating sedentary lifestyle, has rapidly transformed Homo sapiens, in less than a century, into an obese species. Even worse, this transformation may already be impacting future generations via deleterious epigenetic programming. Despite the fact that many people maintain a relatively constant energy balance over lifespan, the doubling of the prevalence of obesity and related metabolic diseases in just 30 years, faster than our genome may adapt, arguably implies that evolutionary-conserved homeostatic mechanisms tightly control the energy regulatory system biased toward efficiently defending energy stores over losses. Consequently, understanding the complex processes which promote excess weight gain and its homeostatic defense mechanisms represents a major challenge to 21st century biomedical research. "All disease begins in the gut" (Hippocrates) This quote from our Greek forefather dates back nearly 2,500 years. This is all the more remarkable as he was not far off. Intriguingly, younger medical history - partly coming from the operating theatre shows us: healing may not be so far away. A main interest of our research is better understanding the impact of environmental factors and lifestyle on human physiology (such as circadian rhythms, light-dark cycle, distress, diet, xenobiotics, bariatric surgery etc.) and the sensory mechanisms by which the human body reacts to changes in the environmental conditions. Our particular interest is the interplay between the gut microbiome, the immune system, and their interaction with the central nervous system as perceptive entities of the human organism. While the gut microbiota - the collection of microorganisms residing in the mammalian intestine - has long been considered as a predominantly passive member of the human ecosystem with mostly digestive functions, it is now clear that the microbiota is an integral part of systemic human physiology. The gut microbiota communicates with the central nervous system via the synthesis, processing and systemic release of metabolite signaling molecules to shape our endocrine and metabolic output system through mechanisms of nutrient handling and energy expenditure. Remarkably, many of our environmental influences could be shown to shape the microbial ecology and functional capacity, reasoning to hypothesize that therapeutic modulation of the microbiome might be harnessed to alter patients risk for the manifestation of a specific disease. Our group is studying how environmental impacts regulate the inter-organ signaling network referred to as the microbiota-gut-brain axis, to define how targeted modulation of gut microbial composition and their metabolic activity affects neuronal circuits that govern energy homeostasis and endocrine function through output function on energy intake and its combustion within the adipose tissue. Sitting at the intersection between basic and clinical research, we employ a multi-disciplinary approach combining cell culture, preclinical and clinical models to gain a holistic view on the molecular and cellular processes of the gut-brain axis that shape endocrine and metabolic health. Our lab is using an integrated physiology approach that relies greatly upon detailed in vivo phenotyping of genetically and non-genetically modified organisms. Together with Systems Biology approach, coupling integrated multi-omics analyses with cutting-edge molecular cell biology tools, enables us to identify metabolic networks relevant for metabolic disease and translate findings into potential novel therapeutic targets for obesity and related endocrine diseases.
PRICES AND AWARDSο Else Kröner-Exzellenzstipendium (2018) ο Schoeller-Junkmann Prize (2018) ο Novartis Prize (2012) ο Pfizer Young Investigator Award (2011) ο DFG Research Scholarship (2010-2012) FUNDING
OPEN POSITIONSWe are happy to announce the opening of the following research positions: SELECTED PUBLICATIONS:Chen, J., Haase, N., Haange, S.-B., Sucher, R., Münzker, J., Jäger, E., Schischke, K., Seyfried, F., Bergen,
M. von, Hankir, M. K., Krügel, U., and Fenske, W. K. (2021) Fries, C. M., Bae, Y. J., Rayes, N., Sandner, B., Isermann, B., Stumvoll, M., Fagotto, V., Reincke, M.,
Bidlingmaier, M., Mandy, V., Kratzsch, J., and Fenske, W. K. (2020) Palladino, V. S., Chiocchetti, A. G., Frank, L., Haslinger, D., McNeill, R., Radtke, F., Till, A., Haupt, S.,
Brüstle, O., Günther, K., Edenhofer, F., Hoffmann, P., Reif, A., and Kittel-Schneider, S. (2020) Haange, S.-B., Jehmlich, N., Krügel, U., Hintschich, C., Wehrmann, D., Hankir, M., Seyfried, F., Froment, J.,
Hübschmann, T., Müller, S., Wissenbach, D. K., Kang, K., Buettner, C., Panagiotou, G., Noll, M., Rolle-Kampczyk,
U., Fenske, W., and Bergen, M. von (2020) Weykopf, B., Haupt, S., Jungverdorben, J., Flitsch, L. J., Hebisch, M., Liu, G.-H., Suzuki, K., Belmonte, J. C. I.,
Peitz, M., Blaess, S., Till, A., and Brüstle, O. (2019) Christ-Crain, M., Bichet, D. G., Fenske, W. K., Goldman, M. B., Rittig, S., Verbalis, J. G., and
Verkman, A. S. (2019) Sheng, C., Jungverdorben, J., Wiethoff, H., Lin, Q., Flitsch, L. J., Eckert, D., Hebisch, M., Fischer, J.,
Kesavan, J., Weykopf, B., Schneider, L., Holtkamp, D., Beck, H., Till, A., Wüllner, U., Ziller, M. J.,
Wagner, W., Peitz, M., and Brüstle, O. (2018) Fenske, W., Refardt, J., Chifu, I., Schnyder, I., Winzeler, B., Drummond, J., Ribeiro-Oliveira, A.,
Drescher, T., Bilz, S., Vogt, D. R., Malzahn, U., Kroiss, M., Christ, E., Henzen, C., Fischli, S., Tönjes, A.,
Mueller, B., Schopohl, J., Flitsch, J., Brabant, G., Fassnacht, M., and Christ-Crain, M. (2018) Fenske, W., Refardt, J., and Christ-Crain, M. (2018) Jungverdorben, J., Till, A., and Brüstle, O. (2017) Hankir, M. K., Kranz, M., Keipert, S., Weiner, J., Andreasen, S. G., Kern, M., Patt, M., Klöting, N.,
Heiker, J. T., Brust, P., Hesse, S., Jastroch, M., and Fenske, W. K. (2017) Hankir, M. K., Patt, M., Patt, J. T. W., Becker, G. A., Rullmann, M., Kranz, M., Deuther-Conrad, W.,
Schischke, K., Seyfried, F., Brust, P., Hesse, S., Sabri, O., Krügel, U., and Fenske, W. K. (2017) Till, A., Saito, R., Merkurjev, D., Liu, J.-J., Syed, G. H., Kolnik, M., Siddiqui, A., Glas, M.,
Scheffler, B., Ideker, T., and Subramani, S. (2015) Hankir, M. K., Seyfried, F., Hintschich, C. A., Diep, T.-A., Kleberg, K., Kranz, M., Deuther-Conrad, W.,
Tellez, L. A., Rullmann, M., Patt, M., Teichert, J., Hesse, S., Sabri, O., Brust, P., Hansen, H. S.,
Araujo, I. E. de, Krügel, U., and Fenske, W. K. (2017) Christ-Crain, M. and Fenske, W. (2016) Hankir, M. K., Cowley, M. A., and Fenske, W. K. (2016) Hankir, M. K., Kranz, M., Gnad, T., Weiner, J., Wagner, S., Deuther-Conrad, W., Bronisch, F., Steinhoff, K.,
Luthardt, J., Klöting, N., Hesse, S., Seibyl, J. P., Sabri, O., Heiker, J. T., Blüher, M., Pfeifer, A.,
Brust, P., and Fenske, W. K. (2016) Börgeson, E., Johnson, A. M. F., Lee, Y. S., Till, A., Syed, G. H., Ali-Shah, S. T., Guiry, P. J., Dalli, J.,
Colas, R. A., Serhan, C. N., Sharma, K., and Godson, C. (2015) |
|
|
|
|
|
Veranstaltungen | Leistungsberichte | Links |
|
|
Studium | Bibliothek | Impressum |
|
| |
Förderverein | Datenschutzerklärung |
|




| © Copyright 2017 - 2024 |
zum Seitenanfang |